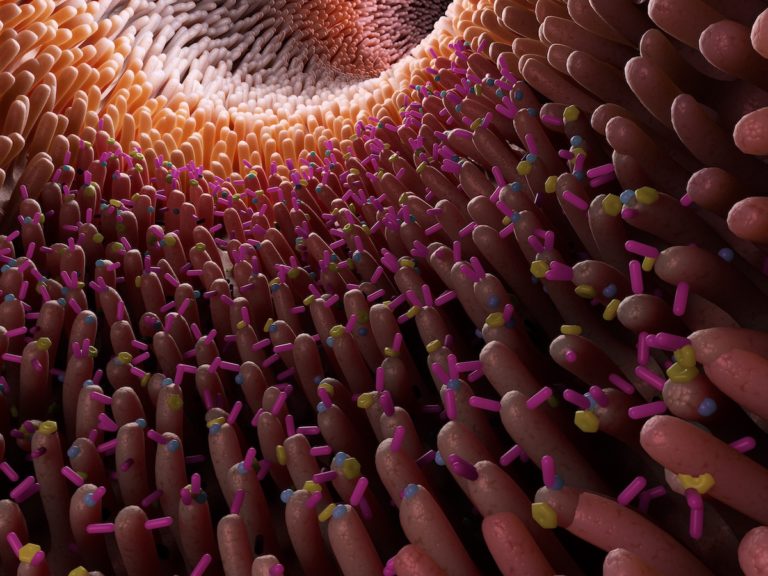
The human microbiome inhabits diverse microbial communities containing various body sites, establishing a symbiotic equilibrium within the host. Throughout our lifespan, these communities undergo changes in function and composition, which are crucial for promoting human health. However, certain perturbations in the microbiome can lead to the deregulation of immune function, metabolism, or the onset of diseases. Observing these perturbations could therefore provide understanding of their impact on our biota, revealing ways to keep a balanced microbiome and protect overall health.
We at 4bases present two sequencing kits that allow the identification of different bacterial and fungal strains in a particular microbiome community. In the case of the MICROBIOME PLUS panel LONG kit, it targets critical genes pivotal for ribosomal RNA function, whilst our MICROBIOME WGS kit employs a whole genome sequencing approach; both kits enabling comprehensive analysis of entire microbiome communities, and identification of the strains contained within them. Both kits utilise the sequencer technology from Oxford Nanopore Technologies, and were tested on bacterial (gut, skin, vagina) and fungal (mycobiome) niches.
The MICROBIOME PLUS panel LONG kit demonstrated high efficacy in identifying various strains within the microbe communities, with over 96% of reads accurately mapped to control strains. Additionally, it correctly estimated the relative abundances of each strain across gut, skin, and vaginal samples. However, when the different strains were too closely related or belonged to similar taxa, the available data sets and tools to map rRNA sequencing results make it hard to discriminate precisely amongst species.
The MICROBIOME WGS kit offered improved classification of closely related strains through whole-genome sequencing, resulting in enhanced precision. This was particularly evident in the mycobiome analysis, where the different taxa (along with their abundance in the community) were identified with remarkable accuracy. These findings highlight the effectiveness of the WGS approach in discriminating between closely related microbial species, providing valuable insights into complex microbiome compositions.
In conclusion, the utilisation of these kits employing Oxford Nanopore sequencers enables the comprehensive analysis of bacterial and fungal strains within diverse microbiome communities, including those of the skin, gut, vagina, and mycobiome. Through precise strain identification, these kits facilitate the evaluation of potential virulence, pathogenicity factors, and antibiotic resistance within these communities. Ultimately, this information serves as a valuable tool in identifying and developing more effective therapies engineered to fight specific microbial strains, thus advancing personalised healthcare approaches.
Learn more about our sequencing kits here.
Mugnai F, Degiorgi A, Giordano S, Favalli V (2024). A clinical ready, long reads solution for sequencing human microbiota.